This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
MECP2 Protein Sequence
Human MECP2 Protein Sequence (isoform 1) Accession Number: NP_004983.1
The human MECP2 protein is 486 amino acids long. Isoform 1 includes exon 2 and has a distinct N terminus (1)
Homologs
|
Pan troglodytes (Chimpanzee) |
Mus musculus (Mouse) |
I used homologene to identify 6 homologs in different organisms. (2) It is important to note that the protein sequences with accession numbers beginning with "XP" (Chimpanzee, Dog, and Cow) are predicted sequences. This resource was very easy to use and each homolog had links to the DNA and Protein sequences in entrez. However, even though results were given back rapidly, they were limited to the 6 most similar homologs. One weakness of homologene is that it doesn't give the E-value. So, I used BLAST to determine the E-value and % Identity for each individual homolog. (3)
I also used BLAST to search for homologs in other organisms, such as Drosophila melanogaster, C. elegans, and Arabidopsis thaliana but found no significant similarities to the Human MECP2 protein sequence. (3)
The homolog results show that the protein sequences are very similar in mammals with the Pan troglodytes homolog being the most similar. The Danio rerio homolog was the most dissimilar and was also the only homolog that wasn't a mammal.
Protein Alignments
|
| ||||||||||||
I used 2 multiple alignment programs, t-coffee (4) and clustalw (5), to compare the protein sequences of the 6 MECP2 homologs. Both programs had similar results but, t-coffee had an additional feature where the amino acids were color coded to portray the similarity of alignments at each specific position. The alignments show that the amino acid sequence is pretty highly conserved, especially in mammals. Danio rerio showed the highest amount of extra amino acids, or missing amino acids. These results agree with the e-values and % Identity found in the BLAST alignments.
Phylogeny
I used 2 programs, tree top (6) and phylogeny.fr (7), to create phylogenetic trees, portraying a visual representation of the evolutionary relationship of each homolog.
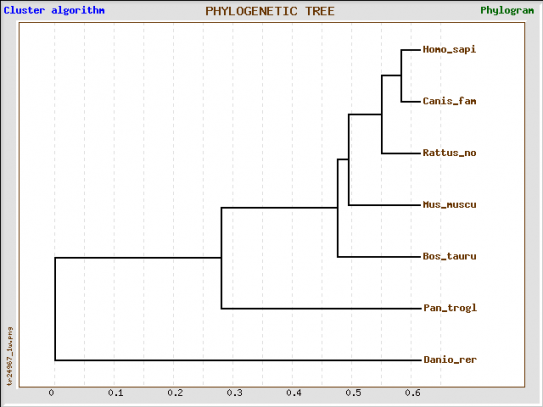
Figure 1: Phylogenetic tree of Human MECP2 protein and 6 homologs produced by Tree top
The phylogenetic tree produced by Tree top (Figure 1) suggests that the Human MECP2 protein is most related to the Dog MECP2 Protein. The Chimpanzee and Zebrafish MECP2 proteins are the most distant from Human MECP2.
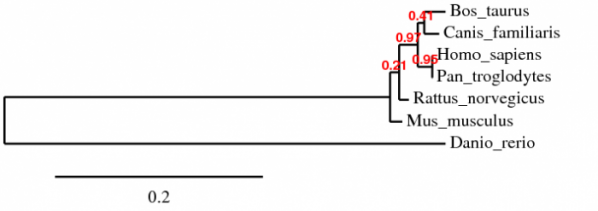
Figure 2: Phylogenetic tree of Human MECP2 protein and 6 homologs produced by phylogeny.fr
The phylogenetic tree produced by phylogeny.fr (above) produced very different results. Human MECP2 was most related to Chimpanzee MECP2. Cow and Dog MECP2 were most related to each other and Zebrafish MECP2 was the most distant from the rest of the homologs.
These results show the variability in using programs to create phylogenetic trees. Since the majority of my homologs had over 90% identical amino acid sequences, it may have been more difficult to get an accurate phylogenetic tree.
Protein Domains
I used SMART (8) and Pfam (9) to determine the domains found in the MECP2 protein sequence. Both programs returned similar results. The most significant domain was the Methyl CpG Binding Domain (MBD) domain with an e-value of 7.343-36*. The MBD domain binds to DNA that has at least one methylated CpG. This is associated with chromatin alteration and gene silencing (10).
Both programs also identified two AT hook motifs with e-values of 2.19e+00* and 1.20e+02*, but they were less significant. AT hook motifs bind to DNA sequences with A/T rich regions. SMART also identified several low complexity regions.
An additional insignificant domain that Pfam identified was SR-25, a Nuclear RNA-splicing-associated protein, with an e-value of 0.95*. This protein is expressed in almost all tissues and contains several nuclear localization signals, suggesting that it is a nuclear protein involved in RNA splicing (11).
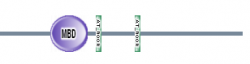
SMART had an additional feature where you could perform a batch analysis. I used this feature with my 7 homologs and found the results seen in figures 3 and 4 below. All six of the homologs I am studying contained the MBD domain with a high e-value. Danio rerio was the only organism that did not contain the AT hook motifs. These results suggest that that the MBD domain is highly conserved in MECP2 proteins of various organisms.
*E-values are taken from SMART with the exception of the SR-25 e-value, which was taken from Pfam. The MBD and AT Hook e-values in Pfam were only slightly different and showed the same general trend in terms of significance.
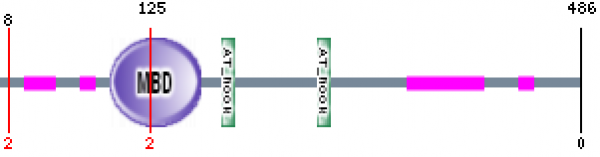
Figure 3: Map of Human MECP2 protein using SMART. The Human MECP2 protein contains an MBD domain, 2 AT hook motifs, and several low complexity regions. (8)
The MBD domain binds DNA at methylated CpG sites and silences genes.
The AT hook domain also binds DNA with a preference for A/T rich regions.
Domains of other Homologs
Figure 4: Map of domains in MECP2 homologs (8).
I was very impressed by the ease of use and the completeness of information provided by both Pfam and SMART. They both gave similar basic information (doming name, location on sequence, and e-value) by simply putting in the protein sequence. Both Pfam and SMART also have links to descriptions of the domains. SMART has some additional features that are particularly useful, such as a schematic of the proteins MECP2 interacts with and a map of the locations of the domains on the protein sequence. MECP2 also has a feature where you can search for domains in multiple sequences as once. This is very useful when comparing homologs. There are also several other links in each domain page in SMART where you can find information, such as what other species have a specific domain, relevant literature on the domain, how variants in the domain result in disease, metabolic pathways, and 3D structures.
Chemical Genetics
I tried several of the websites dealing with chemical interactions with MECP2. Neither ChemBank, PubChem, Drug Bank, or the Human Metabolome Database returned any results.
Protein Data Using Uniprot
Uniprot is a resource with protein sequences and functional information. It is also a good resource for finding primary literature on certain proteins. Some proteins are better annotated than others. Human MECP2 was very well annotated and I was able to find a lot of useful information.
MECP2 Uniprot Accession Number: P51608
Isoforms: Isoform A (canonical sequence) and Isoform B
Isoform B expression is 10 times higher than Isoform A in the brain.
Difference between Isoform A and B: 1-9: MVAGMLGLR → MAAAAAAAPSGGGGGGEEERL
Subcellular Localization
MECP2 is localized to the nucleus. This makes sense, since it functions as a transcriptional repressor.
Gene Ontology
Biological process negative regulation of transcription from RNA polymerase II promoter
Cellular component mitochondrion, nucleus
Molecular function double-stranded methylated DNA binding, transcription corepressor activity
For more information, visit the Gene Ontology/Phenotypes page.
Protein-Protein Interactions
SMARCA2 - a transcriptional coactivator cooperating with nuclear hormone receptors to potentiate transcriptional activation. This interaction makes sense because MECP2 is involved with transcriptional repression.
Protein Domains
Uniprot confirms the data I retrieved from SMART indicating that there is one MBD domain and two AT hook domains.
Secondary Structure
Figure 5: Map of secondary structure for MECP2. The protein has one helix, two strands, and two turns. All of these structures are within the MBD domain. (12)
Natural Variations
Uniprot indicates about 30 natural variants in the MECP2 sequence that are related to Rett Syndrome. These single amino acid variants span the entire protein sequence.
Post Translational Modification
MECP2 is phosphorylated on Ser-423 in the brain upon synaptic activity. This attenuates its repressor activity and seems to regulate dendritic growth and spine maturation.
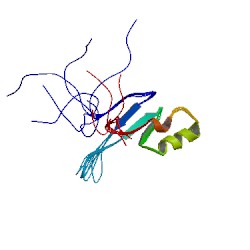
3D Structure
|
Figure 5: (left) 3D Structure of MECP2 protein using NMR (13). |
Protein Data Using ExPASy
Expasy gave very different information compared to uniprot. The information was much more focused on the chemical and molecular properties of the protein.
Molecular weight: 52440.6
Theoretical pI: 9.95
Total number of negatively charged residues (Asp + Glu): 62
Total number of positively charged residues (Arg + Lys): 99
Instability index:
The instability index (II) is computed to be 61.06
This classifies the protein as unstable.
Aliphatic index: 52.59
Grand average of hydropathicity (GRAVY): -1.078
Trypsin cuts my protein in 90 different cleavage sites.
Both Uniprot and ExPASy provided vast amounts of information about my protein. The information was well organized and easy to navigate. Both sites also linked out to several other useful websites.
Protein Interaction Network
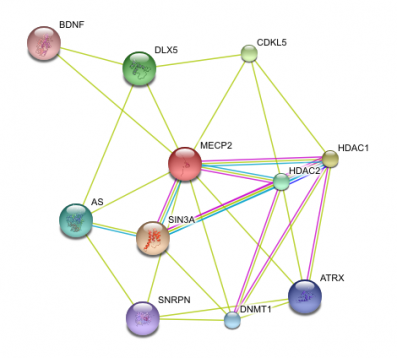
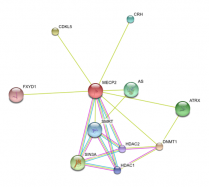
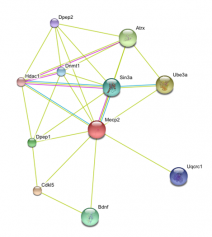
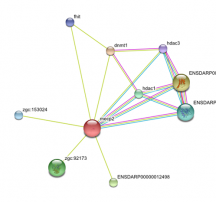
I used String to identify other proteins that MECP2 interacts with. The proteins in this network all have a score of at least 0.959.
SIN3A, HDAC1 and HDAC 2 are all part of the histone deacetylase complex, which leads to chromatin compaction and transcriptional repression (16). This data supports the idea that MECP2 represses transcription.
BDNF is a member of the nerve growth factor family and is necessary for survival of striatal neurons in the brain. It may also be involved in regulating responses to stress. It is proposed to be a major regulator of synaptic transmission and plasticity at adult synapses in many regions of the central nervous system (16). This interaction supports MECP2s role in regulating genes that are involved with neuron function and development.
It is also interesting that MECP2 interacts with CDKL5, a member of the Ser/Thr kinase family (16). Mutations in CDKL5 lead to a rare form of autism that is characterized by early onset and more frequent seizures.
|
Pan troglodytes |
String was very easy to use and gave a nice visual of how different proteins interact with MECP2. It gave clear information as to the evidence that supports these interactions and the confidence we can have that these interactions are true. String also allows users to increase or decrease the number of interacting proteins in the network.
References
1. Entrez - http://www.ncbi.nlm.nih.gov/sites/gquery
2. Homologene - http://www.ncbi.nlm.nih.gov/homologene
3. BLAST (protein) - http://blast.ncbi.nlm.nih.gov/Blast.cgi
4. TCOFFEE - http://www.phylogeny.fr/version2_cgi/one_task.cgi?task_type=tcoffee
5. ClustalW - http://www.phylogeny.fr/version2_cgi/one_task.cgi?task_type=clustalw
6. Tree Top - http://www.genebee.msu.su/services/phtree_reduced.html
7. Phylogeny.fr - http://www.phylogeny.fr/
8. Pfam - http://pfam.sanger.ac.uk/
9. SMART - http://smart.embl-heidelberg.de/
10. Nan X, Meehan RR, Bird A; , Nucleic Acids Res 1993;21:4886-4892.: Dissection of the methyl-CpG binding domain from the chromosomal protein MeCP2. PUBMED:8177735
11. Sasahara K, Yamaoka T, Moritani M, Tanaka M, Iwahana H, Yoshimoto K, Miyagawa J, Kuroda Y, Itakura M; , Biochem Biophys Res Commun. 2000;269:444-450.: Molecular cloning and expression analysis of a putative nuclear protein, SR-25. PUBMED:10708573
12. Uniprot - http://www.uniprot.org/
13. Protein Data Bank - http://www.pdb.org/pdb/cgi/explore.cgi?pdbId=1QK9
14. ExPASy - http://us.expasy.org/tools/#primary
15. String - http://www.weebly.com/weebly/main.php#
16. GeneCards - http://www.genecards.org/
Jessica Connor
[email protected]
Last Updated: 5/11/09